Reading and Writing Data
readr and haven
2025-08-09
readr

| Function | Reads |
|---|---|
read_csv() |
Comma separated values |
read_csv2() |
Semi-colon separate values |
read_delim() |
General delimited files |
read_fwf() |
Fixed width files |
read_log() |
Apache log files |
read_table() |
Space separated files |
read_tsv() |
Tab delimited values |
Importing Data
R functions
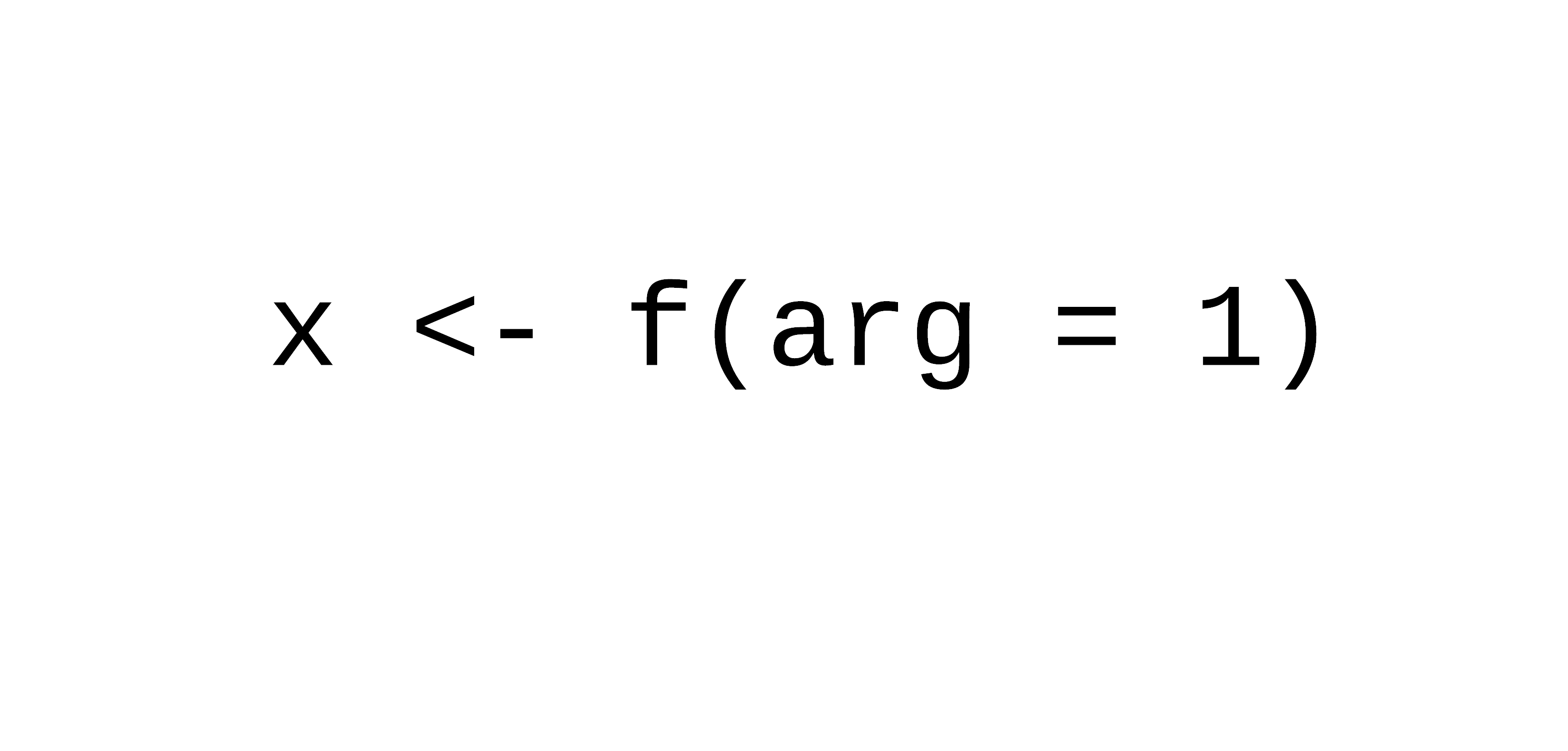
R functions
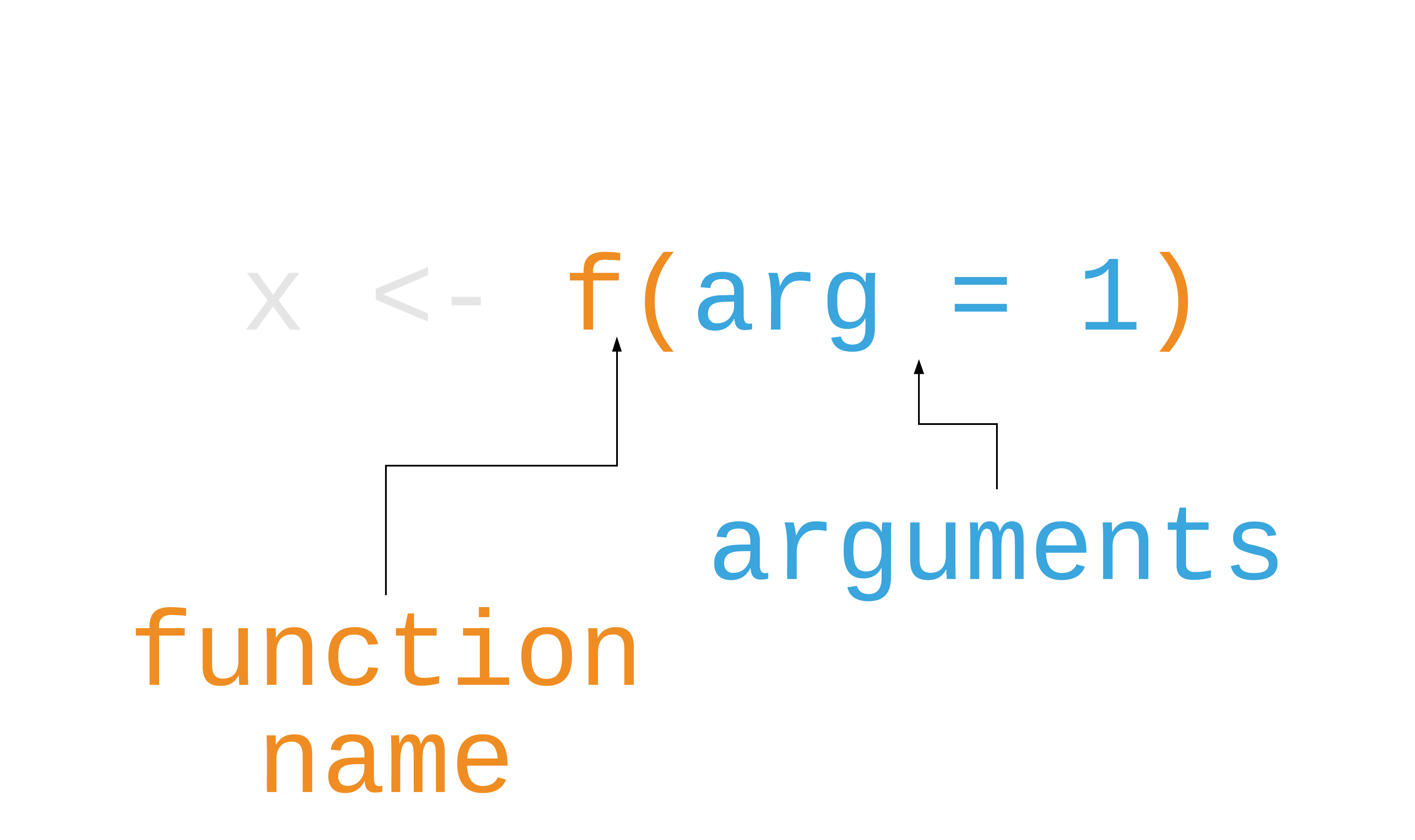
R functions

Your Turn 1
Find diabetes.csv on your computer. Then read it into an object. Then view the results.
Your Turn 1
Find diabetes.csv on your computer. Then read it into an object. Then view the results.

# A tibble: 403 × 19
id chol stab.glu hdl ratio glyhb location age
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <chr> <dbl>
1 1000 203 82 56 3.60 4.31 Buckingham 46
2 1001 165 97 24 6.90 4.44 Buckingham 29
3 1002 228 92 37 6.20 4.64 Buckingham 58
4 1003 78 93 12 6.5 4.63 Buckingham 67
5 1005 249 90 28 8.90 7.72 Buckingham 64
6 1008 248 94 69 3.60 4.81 Buckingham 34
7 1011 195 92 41 4.80 4.84 Buckingham 30
8 1015 227 75 44 5.20 3.94 Buckingham 37
9 1016 177 87 49 3.60 4.84 Buckingham 45
10 1022 263 89 40 6.60 5.78 Buckingham 55
# ℹ 393 more rows
# ℹ 11 more variables: gender <chr>, height <dbl>,
# weight <dbl>, frame <chr>, bp.1s <dbl>, bp.1d <dbl>, …Tibbles
data.frames are the basic form of rectangular data in R (columns of variables, rows of observations)
read_csv() reads the data into a tibble, a modern version of the data frame.
a tibble is a data frame
Missing values
It’s common to use codes for missing values (-99, 9999)
The na option can change these values to NA
Parsing data types
The read functions in readr try to guess each data type, but sometimes it’s wrong
To tell readr how to parse the columns, add the argument col_types to read_csv()
Parsing data types
Or use a string for each variable type: col_type = "cci"
Parsing data types
Or use a string for each variable type: col_type = “cci”
| letter | type |
|---|---|
c |
character |
i |
integer |
n |
number |
d |
double |
l |
logical |
D |
date |
T |
date time |
t |
time |
? |
guess the type |
_ or - |
skip the column |
Your Turn 2
Set the 4 column types to be: integer, double, character, and unknown (guess)
Your Turn 2
Set the 4 column types to be: integer, double, character, and unknown (guess)
haven

| Function | Software |
|---|---|
read_sas() |
SAS |
read_xpt() |
SAS |
read_spss() |
SPSS |
read_sav() |
SPSS |
read_por() |
SPSS |
read_stata() |
Stata |
read_dta() |
Stata |
Heads up!
haven is not a core member of the tidyverse. That means you need to load it with library(haven).
Your Turn 3
There are several versions of the diabetes file besides CSV. Pick a file format you or your colleagues use and import them using the corresponding function from haven.
Your Turn 3
Your Turn 3
# A tibble: 403 × 19
id chol stab_glu hdl ratio glyhb location age
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <chr> <dbl>
1 1000 203 82 56 3.60 4.31 Buckingham 46
2 1001 165 97 24 6.90 4.44 Buckingham 29
3 1002 228 92 37 6.20 4.64 Buckingham 58
4 1003 78 93 12 6.5 4.63 Buckingham 67
5 1005 249 90 28 8.90 7.72 Buckingham 64
6 1008 248 94 69 3.60 4.81 Buckingham 34
7 1011 195 92 41 4.80 4.84 Buckingham 30
8 1015 227 75 44 5.20 3.94 Buckingham 37
9 1016 177 87 49 3.60 4.84 Buckingham 45
10 1022 263 89 40 6.60 5.78 Buckingham 55
# ℹ 393 more rows
# ℹ 11 more variables: gender <chr>, height <dbl>,
# weight <dbl>, frame <chr>, bp_1s <dbl>, bp_1d <dbl>, …Writing data
| Function | Writes |
|---|---|
write_csv() |
Comma separated values |
write_excel_csv() |
CSV that you plan to open in Excel |
write_delim() |
General delimited files |
write_file() |
A single string, written as is |
write_lines() |
A vector of strings, one string per line |
write_tsv() |
Tab delimited values |
write_rds() |
A data type used by R to save objects |
write_xpt() |
SAS transport format, .xpt |
write_sas() |
SAS .sas7bdat files (experimental) |
write_sav() |
SPSS .sav files |
write_stata() |
Stata .dta files |